Biotechnological
Communication
Biosci. Biotech. Res. Comm. 9(3): 523-529 (2016)
Classi cation of cancerous and non-cancerous tissues
of serial analysis of gene expression data through
various classi ers
Gotam S. Lalotra
1
and R.S.Thakur
2
Department of Computer Applications, Maulana Azad National Institute of Technology Bhopal,
India-462003
ABSTRACT
Cancer can be cured if detected early and it can be detected by the expression level analyzed in the suspected tissues.
Serial Analysis of Gene Expression (SAGE) is a gene expression technique used to analyze the genes on the basis of
expression level of the genes. The Libraries of SAGE data contained very large number of genes, considering all genes
for classifying is very tedious task and is not wise thing to do. The preprocessing of the SAGE data is performed to
remove the irrelevant genes by comparing the expression level of genes in normal and cancerous libraries, and the
further analysis of the dataset is done considering the reduced genes. This paper compares classi cation techniques
for classifying the cancerous and non-cancerous tissues of human brain. The Naive Bayes (NB), Linear Discriminant
Analyzer (LDA), Decision Table (DT), Support Vectors Machine (SVM) and K Nearest Neighbor (KNN) classi ers have
been implemented for the analysis of SAGE data. WEKA (The Waikato Environment for Knowledge Analysis) open
source software which consists of a collection of machine learning algorithms for data mining used for analysis. The
results obtained reveal that the K-Nearest Neighbor (KNN) and Linear Discriminant Analyzer (LDA) have given better
performance over other classi es in most of the performance measures except few. Different errors measures have
also been studied in this paper for SAGE data of human brain tissues. The KNN and LDA both have given signi cant
improvement over other classi ers.
KEY WORDS: CLASSIFICATION, SAGE, DIMENSIONAL, ERROR, PERFORMANCE
523
ARTICLE INFORMATION:
*Corresponding Author: singh.gotam@gmail.com,
ramthakur2000@yahoo.com
Received 11
th
Aug, 2016
Accepted after revision 15
th
Sep, 2016
BBRC Print ISSN: 0974-6455
Online ISSN: 2321-4007
Thomson Reuters ISI ESC and Crossref Indexed Journal
NAAS Journal Score 2015: 3.48 Cosmos IF : 4.006
© A Society of Science and Nature Publication, 2016. All rights
reserved.
Online Contents Available at: http//www.bbrc.in/
524 SERIAL ANALYSIS OF GENE EXPRESSION DATA THROUGH VARIOUS CLASSIFIERS BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
Gotam Lalotra and R. S. Thakur
Table 1: Sample SAGE data output
Tag CCAAAACCCA ACAAGATTCC ACCAATTCTA GCCCTCTGAA ACCCTAGGAG
Frequency 27 8 56 90 389
INTRODUCTION
Machine learning is the process of learning structure
from data, there are various machine learning techniques
being implemented to learn from the data. Classi cation
is a data mining technique used to predict group mem-
bership for data instances from instances described by a
set of attributes and a class label data mining remains
the hope for revealing patterns that underlie it (Witten
et al., 2011; Li et al., 2016 and Kumar et al., 2016).
There are some basic techniques for data mining like
classi cation, clustering, association rule mining (Piz-
zuti et al., 2003; Marr, 1981; Wong et al., 2008). Various
state of the art classi cation techniques like Naïve Bayes
(Becker et al., 2001), LDA (Quinlan, 1993), SVM (Cortes
et al., 1995; Burges et al., 1998; Han et al. 2012; Cun-
ningham et al. 2007), KNN (Han et al. 2012) and Decision
Table (DT) has been used for analysis of data.
This paper focuses on the study of the SAGE data of
human brain tissues, which is based on the gene expres-
sion techniques for analysis of genes. SAGE data sets
were collected from SAGE libraries from http://www.
ncbi.nlm.nih.gov/projects/SAGEThe classi cation data
were classi ed into one of the prede ned classes and
hence from the machine learning perspective it is a
supervised learning technique. The Gene expression data
is an example of presenting a large number of features
(genes), most of the features are irrelevant to the de ni-
tion of the problem which consequently could degrade
the classi cation process signi cantly while performing
analysis (Banka et al. 2015). This paper primarily focuses
on experimentally evaluating different methods for clas-
sifying cancerous and non- cancerous tissues.
DATASET PREPARATION
Dataset contains 10 Cancerous and 4 normal libraries,
these datasets are represented in the form of Table 1
containing tag and frequency. These libraries in the form
Tag, frequency1, frequency2, frequency3, frequency14
were combined.
ALGORITHM FOR PREPROCESSING
Step 1: The maximum frequency (maxf) and minimum
frequency (minf) of each gene in the normal
libraries was calculated.
Step 2: The frequency of each gene was compared in the
cancerous libraries with the maximum and the
minimum frequency of normal libraries.
Step 3: Let a
ij
is the frequency of gene j in library i.
1. If (a
ij
> maxf) or (a
ij
< minf)
2. Change frequency value to 1
3. And 0 otherwise
Step 4: 1 shows the differently expressed genes in the
tumor tissue and 0 means no change in the
expression level.
Step 5: Records corresponds to ambiguous tags (genes
which show over expression in some cancer tis-
sues and under expression in some other cancer
tissues) are removed.
The above steps were used for preprocessing on dataset
matrix (14×65454) and have been reduced into matrix
size (14×1898).
RESULTS AND DISCUSSION
The comparison was conducted using the WEKA (The
Waikato Environment for Knowledge Analysis) open
source software which consists of a collection of machine
learning algorithms for data mining. Different classi ers
used for evaluation of the cancerous and non- cancerous
tissues are discussed below in Table 2. The Performance
of the Classi er is discussed in Table 3.
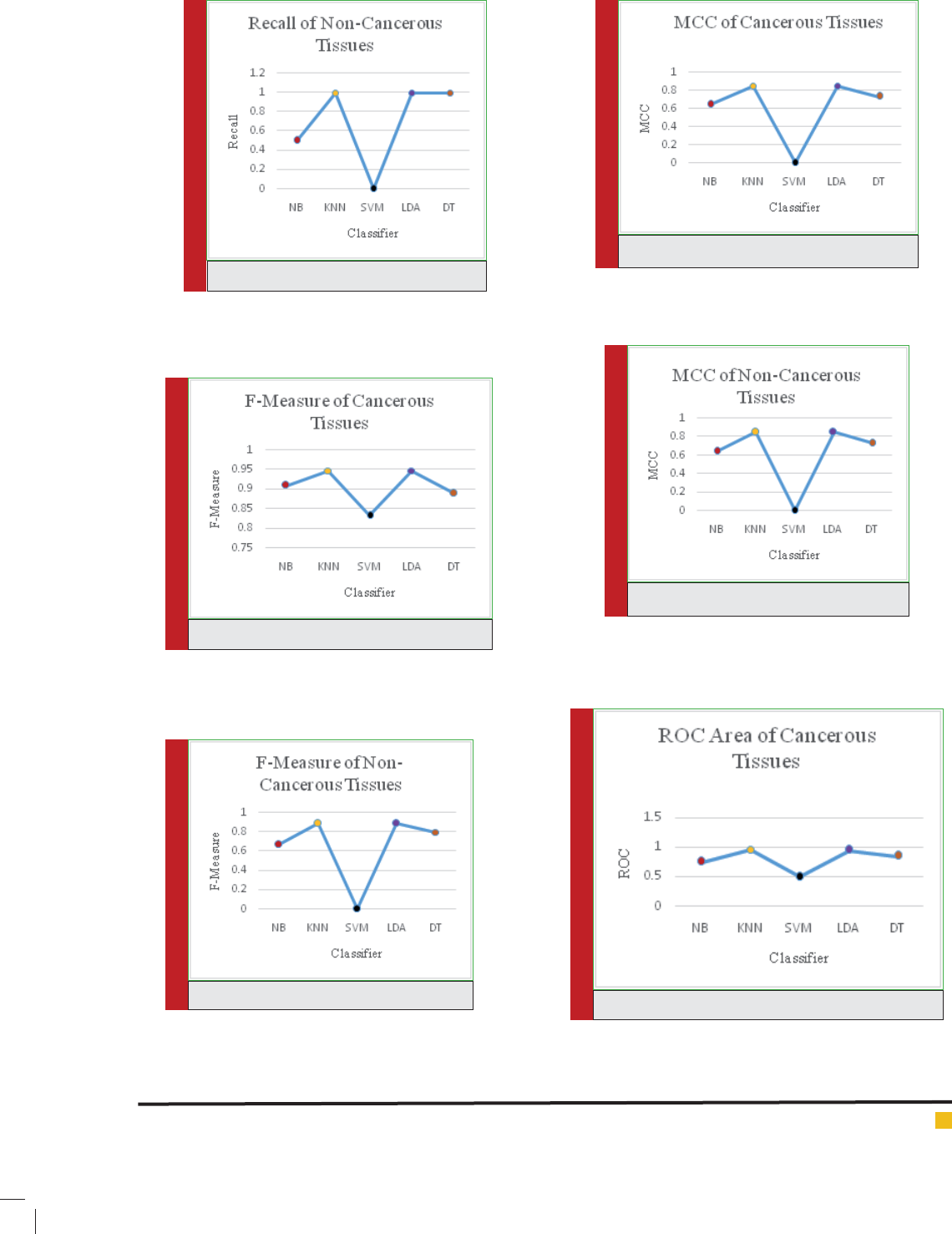
It has been observed that the different classi cation
measures have been calculated and compared for can-
cerous and non-cancerous tissues of human brain. The
measures like True Positive (TP) rate, False Positive (FP)
rate, Precision, Recall, F-Measure, Mathews Correlation
Coef cient (MCC), Receiver Operating Characteristic
(ROC) Area and Precision Recall Curve (PRC) Area have
been used. The all classi ers have performed well after
reducing the number of genes from 65454 to 1898 and
the analysis is performed on the 1898 genes which is
a signi cant improvement in reducing the number of
features but, it can be revealed from the results that
the K-Nearest Neighbor (KNN) and Linear Discriminant
Analyzer (LDA) have outperformed the other classi es in
most of the performance measures.
Discriminant analyzer technique by (Li et al. 2016)
has been proposed to enhance the classi cation accu-
racy. Nearest neighbor classi er requires large memory
and time (Kumar et al. 2016) but, with our algorithm
for preprocessing the dataset has signi cantly reduced
for analysis purpose. A variant of LDA is introduced by
(Bacchus et al. 2013) where LDA has performed better
than SVM and KNN.
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS SERIAL ANALYSIS OF GENE EXPRESSION DATA THROUGH VARIOUS CLASSIFIERS 525
Gotam Lalotra and R. S. Thakur
Table 2: Classi ers and their principle
S. No Name of Classi er Year Principle of Classi er
1 Naive Bayes Classi ers are in use since 1950 Bayesian theorem, it is used when the data is high
dimensional.
2 KNN M. Cover and P. E. Hart in 1967 Closest neighbor whose class is already known.
3 SVM SVM was introduced by Vapnik in
1995 (Wong et al., 2008).
SVM is based on statistical learning theory and
structural risk minimization principal with the aim
of determining location of decision boundaries also
known as hyperplane.
4 LDA LDA was introduced by Fishers
in 1936.
Compute thed-dimensional mean vectors for
the different classes from the dataset, calculate
the scatter matrices. Calculate the eigenvectors
(e1, e2,...,ed) and corresponding eigenvalues
(1,2,...,d) for the scatter matrices. Sort the
eigenvectors by decreasing eigenvalues and choosek
eigenvectors with the largest eigenvalues to form
ad×k dimensional matrixW(where every column
represents an eigenvector). Use thisd×keigenvector
matrix to transform the samples onto the new
subspace. This can be summarized by the matrix
multiplication:Y=X×W.
5 DT Decision tables were introduced
during 1960-70.
one of the earliest classi cation models to be
represented graphically because of their easy to
understand structure (Quinlan, 1993). It is rule based
classi er, where Table represents complete set of
conditional expressions, expressions are mutually
exclusive in in a prede ned area.
Table 3: Classi ers and their performances
S. No Name of
Classi er
Performance of Classi er
1. Naive Bayes For our dataset it has Correctly
Classi ed 12 instances that is the
85.7143 % and 2 instance are
incorrectly classi ed which is
14.2857 %
2. KNN For our dataset it has correctly
classi ed instances are 13 which
is 92.8571 % and the incorrectly
classi ed instances 1 that is 7.1429 %
3. SVM SVM classi ed correctly 10 instance
which is 71.4286 % of the total
and the remaining 28.5714 % are
incorrectly Classi ed Instances.
4. LDA Correctly classi ed instances are13
which is 92.8571 % and incorrectly
classi ed instance is 1 that is
7.1429 %. By LDA.
5 DT In our experiment 12 instances are
correctly classi ed which is 85.7143
% and 2 instances are incorrectly
classi ed which accounts for
14.2857 % by DT.
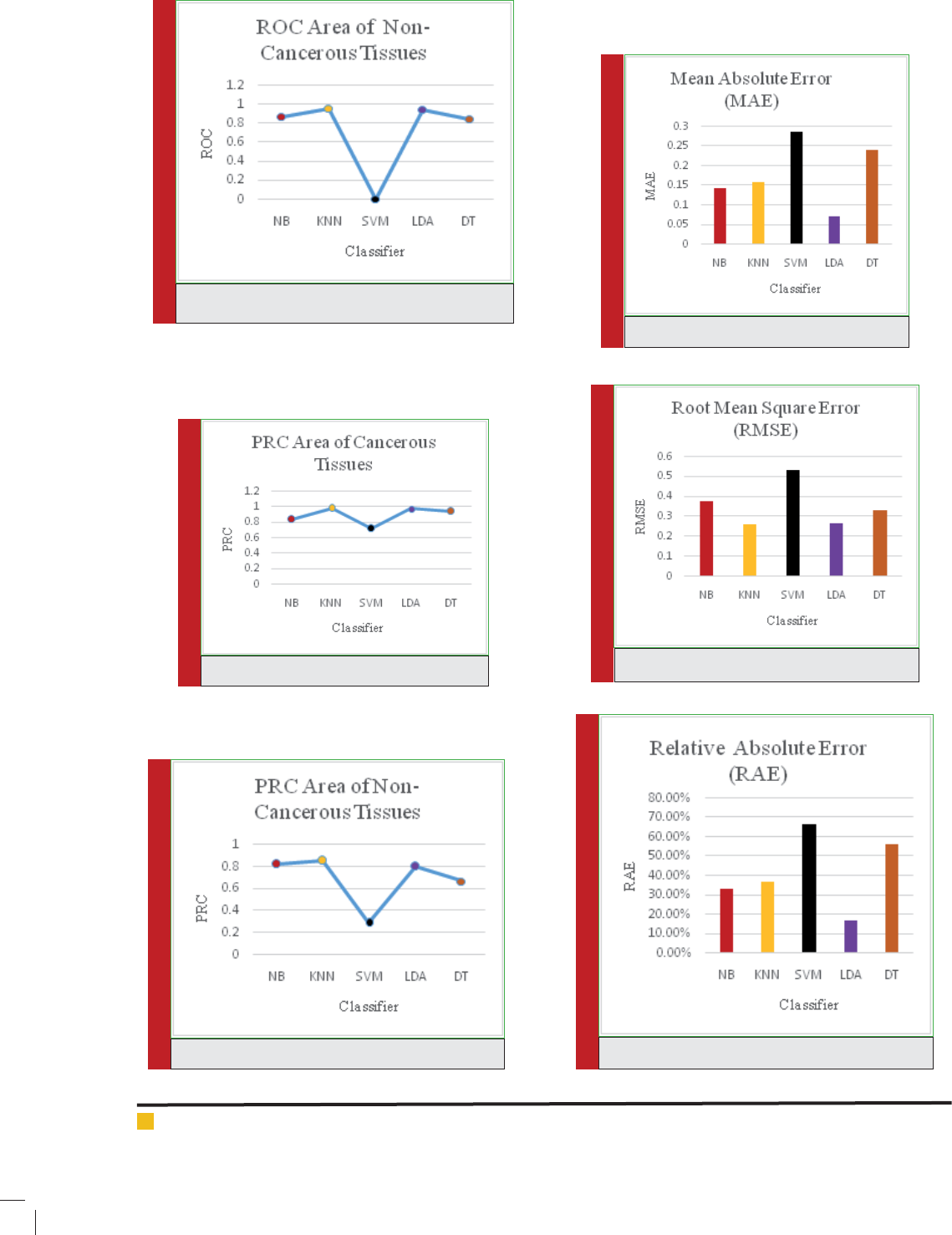
ure Mean Absolute Error (MAE), Root Means Square
Error (RMSE), Relative Absolute Error (RAE) and Root
Relative Square Error (RRSE). These measures also
show that LDA and KNN have performed very well
and the error are less in comparison to other classi ers
used.
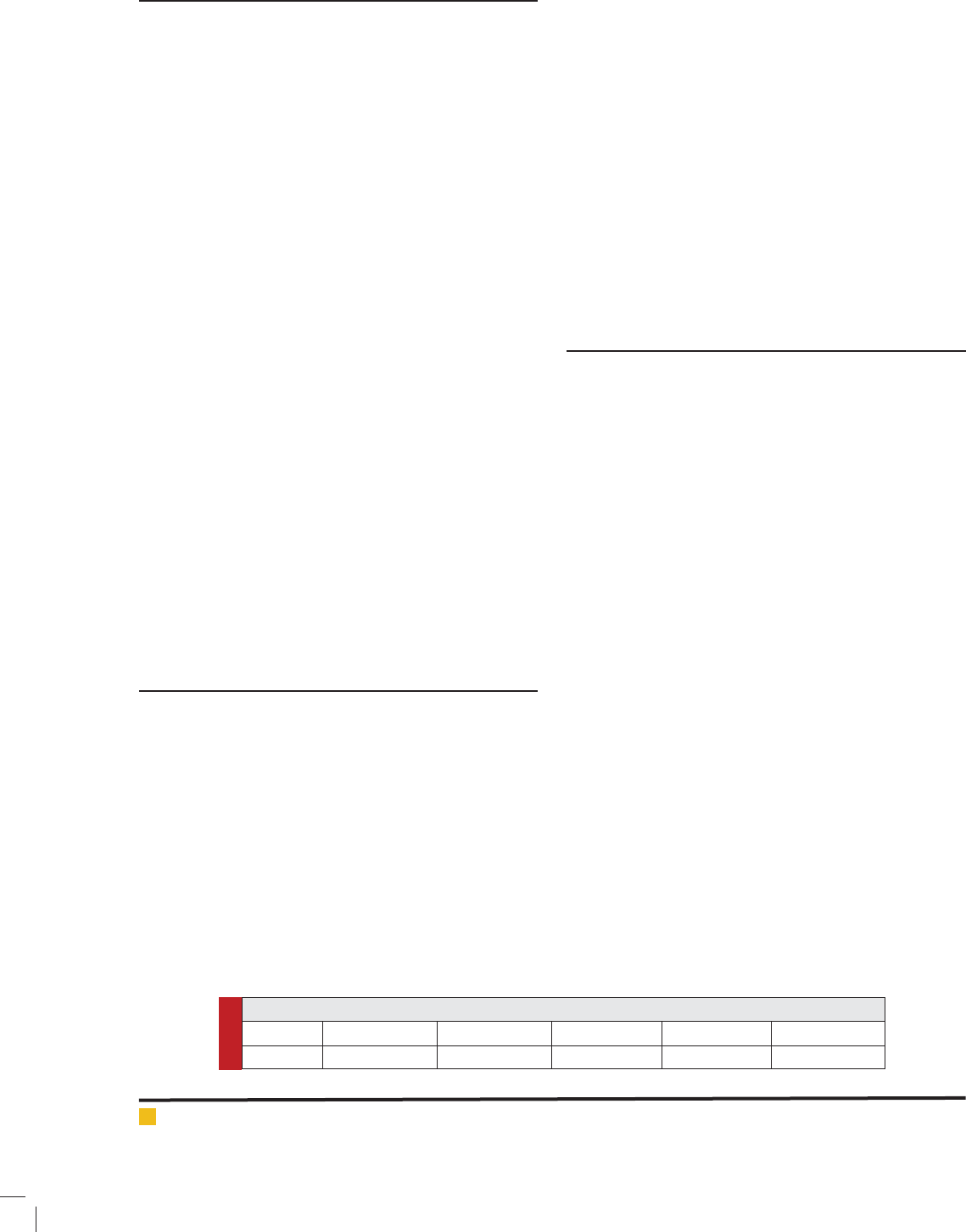
Comparison of Different Performance measures for
Tumorous Tissues and Non-Tumorous Brain Tissues
FIGURE 1.
With the help of preprocessing algorithm, we have
achieved very good results for SAGE dataset as show
in Table 3. The error measures in Figure 17, Figure 18,
Figure 19 and Figure 20 Show the different error meas-
526 SERIAL ANALYSIS OF GENE EXPRESSION DATA THROUGH VARIOUS CLASSIFIERS BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
Gotam Lalotra and R. S. Thakur
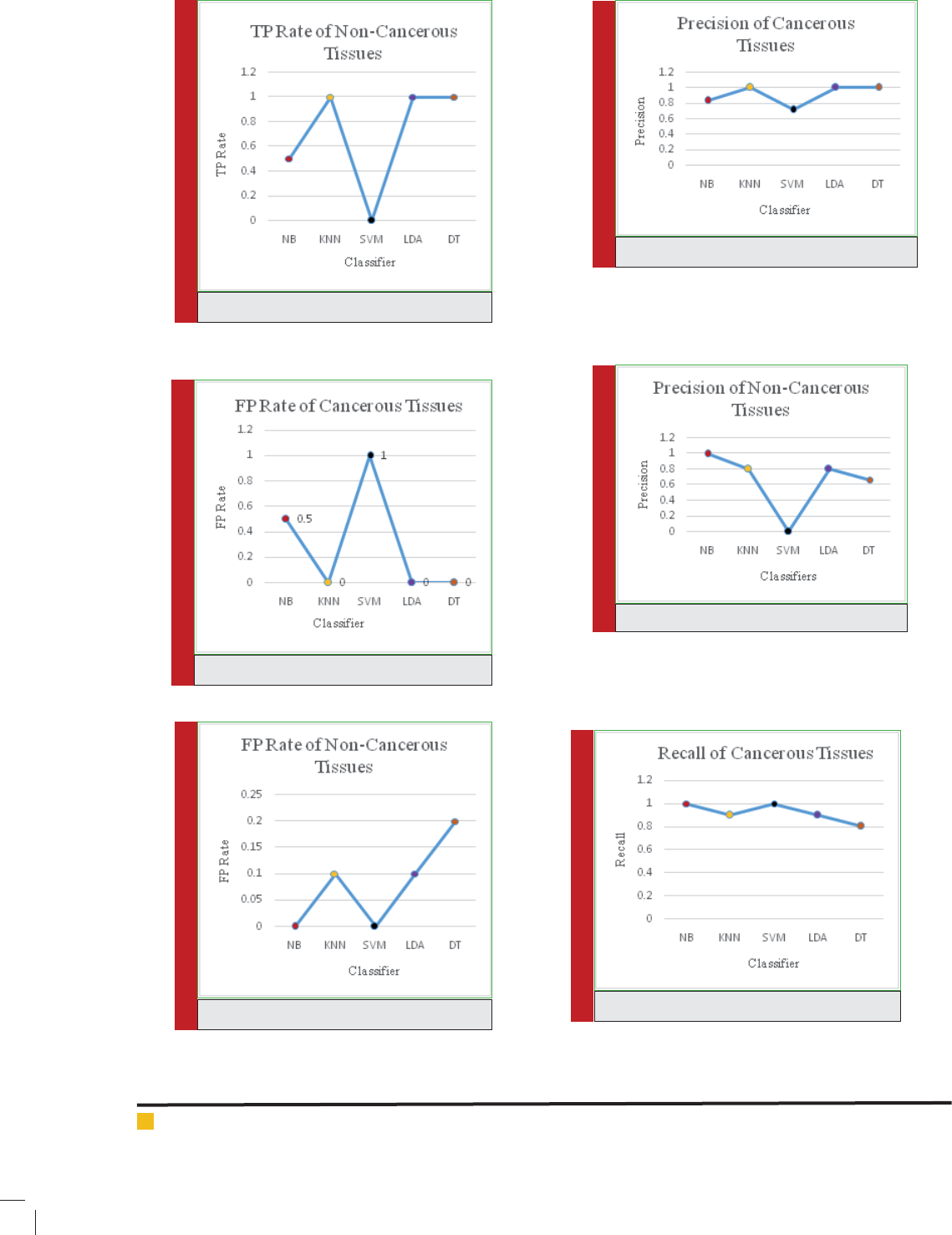
FIGURE 2.
FIGURE 3.
FIGURE 4.
FIGURE 5.
FIGURE 6.
FIGURE 7.
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS SERIAL ANALYSIS OF GENE EXPRESSION DATA THROUGH VARIOUS CLASSIFIERS 527
Gotam Lalotra and R. S. Thakur
FIGURE 10.
FIGURE 8.
FIGURE 9.
FIGURE 11.
FIGURE 12.
FIGURE 13.
528 SERIAL ANALYSIS OF GENE EXPRESSION DATA THROUGH VARIOUS CLASSIFIERS BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
Gotam Lalotra and R. S. Thakur
FIGURE 16.
FIGURE 19.
FIGURE 14.
FIGURE 15.
Error Measures for various Classi ers
FIGURE 18.
FIGURE 17.

Gotam Lalotra and R. S. Thakur
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS SERIAL ANALYSIS OF GENE EXPRESSION DATA THROUGH VARIOUS CLASSIFIERS 529
FIGURE 20.
CONCLUSION
SAGE data has been preprocessed and thereby reduc-
ing the number of features to 1898, samples are clas-
si ed and their performance is compared. The classi-
ers employed here have used statistical approaches
and focused on individual features. Future work can be
enhanced to the study of features in the groups. Imple-
menting the association rule mining techniques along
with soft fuzzy techniques can be of great signi cance
for the reduction in the number of features and perfor-
mance enhancement can be achieved.
ACKNOWLEDGEMENT
This work was supported by research grant from MANIT,
Bhopal, India under Grants in Aid Scheme 2010-11, No.
Dean (R&C)/2010/63 dated 31/08/2010.
REFERENCES
Bacchus A, Biet M, Macaire L, Menach Y L and Tounzi A
(2013). Comparison of Supervised Classification Algorithms
Combined with Feature Extraction and Selection: Application
to a Turbo-generator Rotor Fault Detection, 9th IEEE Interna-
tional Symposium on Diagnostics for Electric Machines, Power
Electronics and Drives (SDEMPED). 558-565.
Banka H, Dara S (2015). A Hamming distance based binary
particle swarm optimization (HDBPSO) algorithm for high
dimensional feature selection, classi cation and validation.
Pattern Recognition Letters. Vol 52, 94–100.
Becker B, Kohavi R, Sommer eld D (2001). Visualizing the
simple Baysian classi er, in: Information Visualization in Data
Mining and Knowledge Discovery. Inc. San Francisco, CA,
USA, Morgan Kaufmann Publishers. 237-249.
Burges C. J.C. (1998). A Tutorial on Support Vector Machines
for Pattern Recognition. Data Mining and Knowledge Discov-
ery 2, 121-167.
Cortes C, Vapnik V (1995). Support vector networks. Machine
Learning, Boston, Kluwer Academic Publishers. Vol 20, 273–
297.
Cunningham P, Delany S. J (2007) k-Nearest Neighbour Classi-
ers Technical Report UCD-CSI.
Han J, Kamber M (2012). Data Mining Concepts and Tech-
niques. Elsevier.
Kumar R.R, Viswanath P, Bindu C.S (2016). Nearest Neighbor
Classifiers: Reducing the Computational Demands, 6th IEEE
International Conference on Advanced Computing. 45-50.
Li L, Du L, Zhang W, Hen H, Wang P (2016). Enhancing infor-
mation discriminant analysis: Feature extraction with linear
statistical model and information-theoretic criteria, Pattern
Recognition Vol. 60,554–570.
Marr J.D (1981). Comparison of several clustering algorithms
for data rate compression of LPC parameters. IEEE Interna-
tional Conference on Acoustics, Speech, and Signal Processing
ICASSP’81. Vol 6, 964-966.
Pizzuti C, Talia D (2003). P-AutoClass: scalable parallel clus-
tering for mining large data setsIEEE Transactions on Knowl-
edge and Data Engineering. Vol 15, Issue 03, 629-641.
Quinlan J.R (1993). C4.5: Programs For Machine Learning,
Morgan Kaufmann Publishers, Inc.
Witten I.H, Frank E, Hall M.A (2011). Data mining practical
machine learning tools and techniques, Morgan Kaufmann
publisher, Burlington.
Wong A.K.C, Li G.C.L (2008). Simultaneous pattern and data
clustering for pattern cluster analysis. IEEE Transactions on
Knowledge and Data Engineering. Vol 20, Issue 07, 911-923.